Scientists are learning about species adaptation by comparing their stem cell-related genes.
The genes regulating pluripotent stem cells in mammals are surprisingly similar across 48 species, Kyoto University researchers report in the journal Genome Biology and Evolution . The study also shows that differences among these ‘gene regulating networks’ might explain how certain features of mammalian pluripotent stem cells have evolved.
Pluripotent stem cells can self-renew and give rise to all other types of cells in the body. Their characteristics are controlled by a network of regulatory genes and molecules, but little is known about how this network has evolved across mammals.
To this end, Ken-ichiro Kamei of Kyoto University’s Institute for Integrated Cell-Material Sciences (iCeMS), with Miho Murayama and Yoshinori Endo of the Wildlife Research Center, compared 134 gene sets belonging to the pluripotency gene regulatory networks of 48 mammalian species.
They found that this network has been highly conserved across species, meaning genetic sequences have remained relatively unchanged over the course of evolution. This high degree of conservation explains why human genetic sequences can reprogram other mammalian tissue cells to turn into pluripotent stem cells. However, since it is also evident that the regulating networks differ across mammals, there might be more efficient combinations of reprogramming factors for each species. Improving techniques for deriving induced pluripotent stem (iPS) cells from mammalian cells, including those from endangered species, could provide a big boost to research and conservation.
“We have been trying to generate induced pluripotent stem cells from various mammalian species, such as the endangered Grévy’s zebra and the bottlenose dolphin,” says Kamei.
Interestingly, the team found relatively high evolutionary changes in genes just downstream of one of the core gene regulatory networks. “This could indicate that mammalian pluripotent stem cells have diversified more than we thought,” says Inoue-Murayama.
The differences between gene regulatory networks in mammalian pluripotent stem cells might also be associated with unique adaptions.
For example, the naked mole rat has been positively selected for a pluripotency regulatory gene that could be involved in giving it its extraordinary longevity and cancer resistance. The gene might also be involved in the development of the extremely sensitive hairs that help them navigate underground.
The researchers also found evidence of positive selection for certain pluripotency gene regulatory network genes involved in the adaptation of large animals, such as the minke whale, the African elephant and the flying fox, to their environments. Surprisingly, these same genes are associated with cancer in other mammals. Since these large animals are known for being relatively resistant to cancer, the researchers suggest that the adaptive alterations these genes underwent in these animals somehow also changed some of their functions, thus giving this group a degree of cancer resistance.
The researchers say the study is among the first to compare the pluripotency gene regulatory networks across major taxa, and could be applicable to evolutional biology studies and for facilitating and improving the generation of induced pluripotent stem cells from new species.
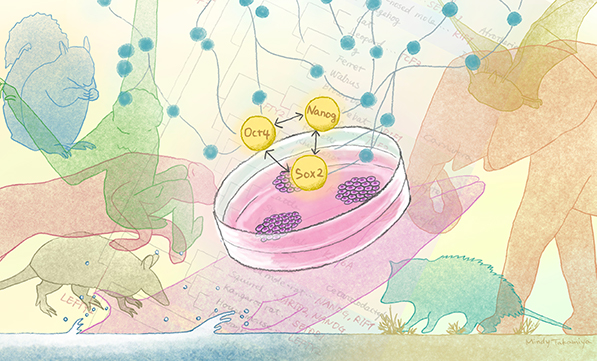
The researchers compared 134 gene sets belonging to the pluripotency gene regulatory networks of 48 mammalian species, and found that this network is highly conserved across species. ((c)Mindy Takamiya/Kyoto University iCeMS - CC BY 4.0)
Paper information
【DOI】 https://doi.org/10.1093/gbe/evaa169
【KURENAI ACCESS URL】 http://hdl.handle.net/2433/254136
Yoshinori Endo, Ken-ichiro Kamei, Miho Inoue-Murayama (2020). Genetic signatures of evolution of the pluripotency gene regulating network across mammals. Genome Biology and Evolution, evaa169.